Page Summary
-
This tutorial demonstrates generating time series data in Earth Engine and visualizing it with the Altair library, using drought and vegetation response as an example.
-
The workflow involves using Earth Engine for accessing and reducing spatiotemporal data, pandas for analysis and organization, and Altair for visualization.
-
Steps for preparing data for visualization include filtering, reducing, formatting as a table, transferring to a Python kernel, converting to a pandas DataFrame, altering the DataFrame, and plotting.
-
Examples show analyzing drought severity (PDSI) using calendar heatmaps and bar charts, vegetation productivity (NDVI) with DOY line charts and distribution plots, and exploring the relationship between drought and productivity with scatter plots.
-
The tutorial also demonstrates preparing and visualizing historical climate data (PRISM) and future climate projections (NEX-DCP30) to speculate on potential impacts of future climate on vegetation.
This tutorial provides methods for generating time series data in Earth Engine and visualizing it with the Altair library using drought and vegetation response as an example.
Topics include:
- Time series region reduction in Earth Engine
- Formatting a table in Earth Engine
- Transferring an Earth Engine table to a Colab Python kernel
- Converting an Earth Engine table to a pandas DataFrame
- Data representation with various Altair chart types
Note that this tutorial uses the Earth Engine Python API in a Colab notebook.
Context
At the heart of this tutorial is the notion of data reduction and the need to transform data into insights to help inform our understanding of Earth processes and human's role in them. It combines a series of technologies, each best suited to a particular task in the data reduction process. Earth Engine is used to access, clean, and reduce large amounts of spatiotemporal data, pandas is used to analyze and organize the results, and Altair is used to visualize the results.
Note: This notebook demonstrates an analysis template and interactive workflow that is appropriate for a certain size of dataset, but there are limitations to interactive computation time and server-to-client data transfer size imposed by Colab and Earth Engine. To analyze even larger datasets, you may need to modify the workflow to export FeatureCollection results from Earth Engine as static assets and then use the static assets to perform the subsequent steps involving Earth Engine table formatting, conversion to pandas DataFrame, and charting with Altair.
Materials
Datasets
Climate
Vegetation proxies
Region of interest
The region of interest for these examples is the Sierra Nevada ecoregion of California. The vegetation grades from mostly ponderosa pine and Douglas-fir at low elevations on the western side, to pines and Sierra juniper on the eastern side, and to fir and other conifers at higher elevations.
General workflow
Preparation of every dataset for visualization follows the same basic steps:
- Filter the dataset (server-side Earth Engine)
- Reduce the data region by a statistic (server-side Earth Engine)
- Format the region reduction into a table (server-side Earth Engine)
- Convert the Earth Engine table to a DataFrame (server-side Earth Engine > client-side Python kernel)
- Alter the DataFrame (client-side pandas)
- Plot the DataFrame (client-side Altair)
The first dataset will walk through each step in detail. Following examples will provide less description, unless there is variation that merits note.
Python setup
Earth Engine API
- Import the Earth Engine library.
- Authenticate access (registration verification and Google account access).
- Initialize the API.
import ee
ee.Authenticate()
ee.Initialize(project='my-project')
Other libraries
Import other libraries used in this notebook.
- pandas: data analysis (including the DataFrame data structure)
- altair: declarative visualization library (used for charting)
- numpy: array-processing package (used for linear regression)
- folium: interactive web map
import pandas as pd
import altair as alt
import numpy as np
import folium
Region reduction function
Reduction of pixels intersecting the region of interest to a statistic will be performed multiple times. Define a reusable function that can perform the task for each dataset. The function accepts arguments such as scale and reduction method to parameterize the operation for each particular analysis.
Note: most of the reduction operations in this tutorial use a large pixel scale so that operations complete quickly. In your own application, set the scale and other parameter arguments as you wish.
def create_reduce_region_function(geometry,
reducer=ee.Reducer.mean(),
scale=1000,
crs='EPSG:4326',
bestEffort=True,
maxPixels=1e13,
tileScale=4):
"""Creates a region reduction function.
Creates a region reduction function intended to be used as the input function
to ee.ImageCollection.map() for reducing pixels intersecting a provided region
to a statistic for each image in a collection. See ee.Image.reduceRegion()
documentation for more details.
Args:
geometry:
An ee.Geometry that defines the region over which to reduce data.
reducer:
Optional; An ee.Reducer that defines the reduction method.
scale:
Optional; A number that defines the nominal scale in meters of the
projection to work in.
crs:
Optional; An ee.Projection or EPSG string ('EPSG:5070') that defines
the projection to work in.
bestEffort:
Optional; A Boolean indicator for whether to use a larger scale if the
geometry contains too many pixels at the given scale for the operation
to succeed.
maxPixels:
Optional; A number specifying the maximum number of pixels to reduce.
tileScale:
Optional; A number representing the scaling factor used to reduce
aggregation tile size; using a larger tileScale (e.g. 2 or 4) may enable
computations that run out of memory with the default.
Returns:
A function that accepts an ee.Image and reduces it by region, according to
the provided arguments.
"""
def reduce_region_function(img):
"""Applies the ee.Image.reduceRegion() method.
Args:
img:
An ee.Image to reduce to a statistic by region.
Returns:
An ee.Feature that contains properties representing the image region
reduction results per band and the image timestamp formatted as
milliseconds from Unix epoch (included to enable time series plotting).
"""
stat = img.reduceRegion(
reducer=reducer,
geometry=geometry,
scale=scale,
crs=crs,
bestEffort=bestEffort,
maxPixels=maxPixels,
tileScale=tileScale)
return ee.Feature(geometry, stat).set({'millis': img.date().millis()})
return reduce_region_function
Formatting
The result of the region reduction function above applied to an ee.ImageCollection produces an ee.FeatureCollection. This data needs to be transferred to the Python kernel, but serialized feature collections are large and awkward to deal with. This step defines a function to convert the feature collection to an ee.Dictionary where the keys are feature property names and values are corresponding lists of property values, which pandas can deal with handily.
- Extract the property values from the
ee.FeatureCollectionas a list of lists stored in anee.DictionaryusingreduceColumns(). - Extract the list of lists from the dictionary.
- Add names to each list by converting to an
ee.Dictionarywhere keys are property names and values are the corresponding value lists.
The returned ee.Dictionary is essentially a table, where keys define columns and list elements define rows.
# Define a function to transfer feature properties to a dictionary.
def fc_to_dict(fc):
prop_names = fc.first().propertyNames()
prop_lists = fc.reduceColumns(
reducer=ee.Reducer.toList().repeat(prop_names.size()),
selectors=prop_names).get('list')
return ee.Dictionary.fromLists(prop_names, prop_lists)
Drought severity
In this section we'll look at a time series of drought severity as a calendar heat map and a bar chart.
Import data
- Load the gridded Palmer Drought Severity Index (PDSI) data as an
ee.ImageCollection. - Load the EPA Level-3 ecoregion boundaries as an
ee.FeatureCollectionand filter it to include only the Sierra Nevada region, which defines the area of interest (AOI).
today = ee.Date(pd.to_datetime('today'))
date_range = ee.DateRange(today.advance(-20, 'years'), today)
pdsi = ee.ImageCollection('GRIDMET/DROUGHT').filterDate(date_range).select('pdsi')
aoi = ee.FeatureCollection('EPA/Ecoregions/2013/L3').filter(
ee.Filter.eq('na_l3name', 'Sierra Nevada')).geometry()
Note: the aoi defined above will be used throughout this tutorial. In your own application, redefine it for your own area of interest.
Reduce data
- Create a region reduction function.
- Map the function over the
pdsiimage collection to reduce each image. - Filter out any resulting features that have null computed values (occurs when all pixels in an AOI are masked).
reduce_pdsi = create_reduce_region_function(
geometry=aoi, reducer=ee.Reducer.mean(), scale=5000, crs='EPSG:3310')
pdsi_stat_fc = ee.FeatureCollection(pdsi.map(reduce_pdsi)).filter(
ee.Filter.notNull(pdsi.first().bandNames()))
STOP:
Optional export
If your process is long-running, you'll want to export the pdsi_stat_fc variable as an asset using a batch task. Wait until the task finishes, import the asset, and continue on. Please see the Developer Guide section on exporting with the Python API.
Export to asset:
task = ee.batch.Export.table.toAsset(
collection=pdsi_stat_fc,
description='pdsi_stat_fc export',
assetId='users/YOUR_USER_NAME/pdsi_stat_fc_ts_vis_with_altair')
# task.start()
Import the asset after the export completes:
# pdsi_stat_fc = ee.FeatureCollection('users/YOUR_USER_NAME/pdsi_stat_fc_ts_vis_with_altair')
* Remove comments (#) to run the above cells.
CONTINUE:
Server to client transfer
The ee.FeatureCollection needs to be converted to a dictionary and transferred to the Python kernel.
- Apply the
fc_to_dictfunction to convert fromee.FeatureCollectiontoee.Dictionary. - Call
getInfo()on theee.Dictionaryto transfer the data client-side.
pdsi_dict = fc_to_dict(pdsi_stat_fc).getInfo()
The result is a Python dictionary. Print a small part to see how it is formatted.
print(type(pdsi_dict), '\n')
for prop in pdsi_dict.keys():
print(prop + ':', pdsi_dict[prop][0:3] + ['...'])
<class 'dict'> millis: [1148536800000, 1148968800000, 1149400800000, '...'] pdsi: [3.516955053228609, 3.485140971128909, 3.4552866333000543, '...'] system:index: ['20060525', '20060530', '20060604', '...']
Convert the Python dictionary to a pandas DataFrame.
pdsi_df = pd.DataFrame(pdsi_dict)
Preview the DataFrame and check the column data types.
display(pdsi_df)
print(pdsi_df.dtypes)
millis int64 pdsi float64 system:index object dtype: object
Add date columns
Add date columns derived from the milliseconds from Unix epoch column. The pandas library provides functions and objects for timestamps and the DataFrame object allows for easy mutation.
Define a function to add date variables to the DataFrame: year, month, day, and day of year (DOY).
# Function to add date variables to DataFrame.
def add_date_info(df):
df['Timestamp'] = pd.to_datetime(df['millis'], unit='ms')
df['Year'] = pd.DatetimeIndex(df['Timestamp']).year
df['Month'] = pd.DatetimeIndex(df['Timestamp']).month
df['Day'] = pd.DatetimeIndex(df['Timestamp']).day
df['DOY'] = pd.DatetimeIndex(df['Timestamp']).dayofyear
return df
Note: the above function for adding date information to a DataFrame will be used throughout this tutorial.
Apply the add_date_info function to the PDSI DataFrame to add date attribute columns, preview the results.
pdsi_df = add_date_info(pdsi_df)
pdsi_df.head(5)
Rename and drop columns
Often it is desirable to rename columns and/or remove unnecessary columns. Do both here and preview the DataFrame.
pdsi_df = pdsi_df.rename(columns={
'pdsi': 'PDSI'
}).drop(columns=['millis', 'system:index'])
pdsi_df.head(5)
Check the data type of each column.
pdsi_df.dtypes
PDSI float64 Timestamp datetime64[ns] Year int32 Month int32 Day int32 DOY int32 dtype: object
At this point the DataFrame is in good shape for charting with Altair.
Calendar heatmap
Chart PDSI data as a calendar heatmap. Set observation year as the x-axis variable, month as y-axis, and PDSI value as color.
Note that Altair features a convenient method for aggregating values within groups while encoding the chart (i.e., no need to create a new DataFrame). The mean aggregate transform is applied here because each month has three PDSI observations (year and month are the grouping factors).
Also note that a tooltip has been added to the chart; hovering over cells reveals the values of the selected variables.
alt.Chart(pdsi_df).mark_rect().encode(
x='Year:O',
y='Month:O',
color=alt.Color(
'mean(PDSI):Q', scale=alt.Scale(scheme='redblue', domain=(-5, 5))),
tooltip=[
alt.Tooltip('Year:O', title='Year'),
alt.Tooltip('Month:O', title='Month'),
alt.Tooltip('mean(PDSI):Q', title='PDSI')
]).properties(width=600, height=300)
The calendar heat map is good for interpretation of relative intra- and inter-annual differences in PDSI. However, since the PDSI variable is represented by color, estimating absolute values and magnitude of difference is difficult.
Bar chart
Chart PDSI time series as a bar chart to more easily interpret absolute values and compare them over time. Here, the observation timestamp is represented on the x-axis and PDSI is represented by both the y-axis and color. Since each PDSI observation has a unique timestamp that can be plotted to the x-axis, there is no need to aggregate PDSI values as in the above chart. A tooltip is added to the chart; hover over the bars to reveal the values for each variable.
alt.Chart(pdsi_df).mark_bar(size=1).encode(
x='Timestamp:T',
y='PDSI:Q',
color=alt.Color(
'PDSI:Q', scale=alt.Scale(scheme='redblue', domain=(-5, 5))),
tooltip=[
alt.Tooltip('Timestamp:T', title='Date'),
alt.Tooltip('PDSI:Q', title='PDSI')
]).properties(width=600, height=300)
This temporal bar chart makes it easier to interpret and compare absolute values of PDSI over time, but relative intra- and inter-annual variability are arguably harder to interpret because the division of year and month is not as distinct as in the calendar heatmap above.
Take note of the extended and severe period of drought from 2012 through 2016. In the next section, we'll look for a vegetation response to this event.
Vegetation productivity
NDVI is a proxy measure of photosynthetic capacity and is used in this tutorial to investigate vegetation response to the 2012-2016 drought identified in the PDSI bar chart above.
MODIS provides an analysis-ready 16-day NDVI composite that is well suited for regional investigation of temporal vegetation dynamics. The following steps reduce and prepare this data for charting in the same manner as the PDSI data above; please refer to previous sections to review details.
Import and reduce
- Load the MODIS NDVI data as an
ee.ImageCollection. - Create a region reduction function.
- Apply the function to all images in the time series.
- Filter out features with null computed values.
ndvi = ee.ImageCollection('MODIS/006/MOD13A2').filterDate(date_range).select('NDVI')
reduce_ndvi = create_reduce_region_function(
geometry=aoi, reducer=ee.Reducer.mean(), scale=1000, crs='EPSG:3310')
ndvi_stat_fc = ee.FeatureCollection(ndvi.map(reduce_ndvi)).filter(
ee.Filter.notNull(ndvi.first().bandNames()))
/tmpfs/src/tf_docs_env/lib/python3.9/site-packages/ee/deprecation.py:214: DeprecationWarning: Attention required for MODIS/006/MOD13A2! You are using a deprecated asset. To make sure your code keeps working, please update it. This dataset has been superseded by MODIS/061/MOD13A2 Learn more: https://developers.google.com/earth-engine/datasets/catalog/MODIS_006_MOD13A2 warnings.warn(warning, category=DeprecationWarning)
STOP:
If your process is long-running, you'll want to export the ndvi_stat_fc variable as an asset using a batch task. Wait until the task finishes, import the asset, and continue on.
Please see the above Optional export section for more details.
CONTINUE:
Prepare DataFrame
- Transfer data from the server to the client.
- Convert the Python dictionary to a pandas DataFrame.
- Preview the DataFrame and check data types.
ndvi_dict = fc_to_dict(ndvi_stat_fc).getInfo()
ndvi_df = pd.DataFrame(ndvi_dict)
display(ndvi_df)
print(ndvi_df.dtypes)
NDVI float64 millis int64 system:index object dtype: object
- Remove the NDVI scaling.
- Add date attribute columns.
- Preview the DataFrame.
ndvi_df['NDVI'] = ndvi_df['NDVI'] / 10000
ndvi_df = add_date_info(ndvi_df)
ndvi_df.head(5)
These NDVI time series data are now ready for plotting.
DOY line chart
Make a day of year (DOY) line chart where each line represents a year of observations. This chart makes it possible to compare the same observation date among years. Use it to compare NDVI values for years during the drought and not.
Day of year is represented on the x-axis and NDVI on the y-axis. Each line represents a year and is distinguished by color. Note that this plot includes a tooltip and has been made interactive so that the axes can be zoomed and panned.
highlight = alt.selection(
type='single', on='mouseover', fields=['Year'], nearest=True)
base = alt.Chart(ndvi_df).encode(
x=alt.X('DOY:Q', scale=alt.Scale(domain=[0, 353], clamp=True)),
y=alt.Y('NDVI:Q', scale=alt.Scale(domain=[0.1, 0.6])),
color=alt.Color('Year:O', scale=alt.Scale(scheme='magma')))
points = base.mark_circle().encode(
opacity=alt.value(0),
tooltip=[
alt.Tooltip('Year:O', title='Year'),
alt.Tooltip('DOY:Q', title='DOY'),
alt.Tooltip('NDVI:Q', title='NDVI')
]).add_selection(highlight)
lines = base.mark_line().encode(
size=alt.condition(~highlight, alt.value(1), alt.value(3)))
(points + lines).properties(width=600, height=350).interactive()
/tmpfs/tmp/ipykernel_17651/3208543205.py:1: AltairDeprecationWarning: Deprecated since `altair=5.0.0`. Use 'selection_point()' or 'selection_interval()' instead. These functions also include more helpful docstrings. highlight = alt.selection( /tmpfs/tmp/ipykernel_17651/3208543205.py:9: AltairDeprecationWarning: Deprecated since `altair=5.0.0`. Use add_params instead. points = base.mark_circle().encode(
The first thing to note is that winter dates (when there is snow in the Sierra Nevada ecoregion) exhibit highly variable inter-annual NDVI, but spring, summer, and fall dates are more consistent. With regard to drought effects on vegetation, summer and fall dates are the most sensitive time. Zooming into observations for the summer/fall days (224-272), you'll notice that many years have a u-shaped pattern where NDVI values decrease and then rise.
Another way to view these data is to plot the distribution of NDVI by DOY represented as an interquartile range envelope and median line. Here, these two charts are defined and then combined in the following snippet.
- Define a base chart.
- Define a line chart for median NDVI (note the use of aggregate median transform grouping by DOY).
- Define a band chart using
'iqr'(interquartile range) to represent NDVI distribution grouping on DOY. - Combine the line and band charts.
base = alt.Chart(ndvi_df).encode(
x=alt.X('DOY:Q', scale=alt.Scale(domain=(150, 340))))
line = base.mark_line().encode(
y=alt.Y('median(NDVI):Q', scale=alt.Scale(domain=(0.47, 0.53))))
band = base.mark_errorband(extent='iqr').encode(
y='NDVI:Q')
(line + band).properties(width=600, height=300).interactive()
The summary statistics for the summer/fall days (224-272) certainly show an NDVI reduction, but there is also variability; some years exhibit greater NDVI reduction than others as suggested by the wide interquartile range during the middle of the summer. Assuming that NDVI reduction is due to water and heat limiting photosynthesis, we can hypothesize that during years of drought, photosynthesis (NDVI) will be lower than non-drought years. We can investigate the relationship between photosynthesis (NDVI) and drought (PDSI) using a scatter plot and linear regression.
Dought and productivity relationship
A scatterplot is a good way to visualize the relationship between two variables. Here, PDSI (drought indicator) will be plotted on the x-axis and NDVI (vegetation productivity) on the y-axis. To achieve this, both variables must exist in the same DataFrame. Each row will be an observation in time and columns will correspond to PDSI and NDVI values. Currently, PDSI and NDVI are in two different DataFrames and need to be merged.
Prepare DataFrames
Before they can be merged, each variable must be reduced to a common temporal observation unit to define correspondence. There are a number of ways to do this and each will define the relationship between PDSI and NDVI differently. Here, our temporal unit will be an annual observation set where NDVI is reduced to the intra-annual minimum from DOY 224 to 272 and PDSI will be the mean from DOY 1 to 272. We are proposing that average drought severity for the first three quarters of a year are related to minimum summer NDVI for a given year.
- Filter the NDVI DataFrame to observations that occur between DOY 224 and 272.
- Reduce the DOY-filtered subset to intra-annual minimum NDVI.
ndvi_doy_range = [224, 272]
ndvi_df_sub = ndvi_df[(ndvi_df['DOY'] >= ndvi_doy_range[0])
& (ndvi_df['DOY'] <= ndvi_doy_range[1])]
ndvi_df_sub = ndvi_df_sub.groupby('Year').min(numeric_only=True)
Note: in your own application you may find that a different DOY range is more suitable, change the ndvi_doy_range as needed.
- Filter the PDSI DataFrame to observations that occur between DOY 1 and 272.
- Reduce the values within a given year to the mean of the observations.
pdsi_doy_range = [1, 272]
pdsi_df_sub = pdsi_df[(pdsi_df['DOY'] >= pdsi_doy_range[0])
& (pdsi_df['DOY'] <= pdsi_doy_range[1])]
pdsi_df_sub = pdsi_df_sub.groupby('Year').mean(numeric_only=True)
Note: in your own application you may find that a different DOY range is more suitable, change the pdsi_doy_range as needed.
- Perform a join on 'Year' to combine the two reduced DataFrames.
- Select only the columns of interest: 'Year', 'NDVI', 'PDSI'.
- Preview the DataFrame.
ndvi_pdsi_df = pd.merge(
ndvi_df_sub, pdsi_df_sub, how='left', on='Year').reset_index()
ndvi_pdsi_df = ndvi_pdsi_df[['Year', 'NDVI', 'PDSI']]
ndvi_pdsi_df.head(5)
NDVI and PDSI are now included in the same DataFrame linked by Year. This format is suitable for determining a linear relationship and drawing a line of best fit through the data.
Including a line of best fit can be a helpful visual aid. Here, a 1D polynomial is fit through the xy point cloud defined by corresponding NDVI and PDSI observations. The resulting fit is added to the DataFrame as a new column 'Fit'.
- Add a line of best fit between PDSI and NDVI by determining the linear relationship and predicting NDVI based on PDSI for each year.
ndvi_pdsi_df['Fit'] = np.poly1d(
np.polyfit(ndvi_pdsi_df['PDSI'], ndvi_pdsi_df['NDVI'], 1))(
ndvi_pdsi_df['PDSI'])
ndvi_pdsi_df.head(5)
Scatter plot
The DataFrame is ready for plotting. Since this chart is to include points and a line of best fit, two charts need to be created, one for the points and one for the line. The results are combined into the final plot.
base = alt.Chart(ndvi_pdsi_df).encode(
x=alt.X('PDSI:Q', scale=alt.Scale(domain=(-5, 5))))
points = base.mark_circle(size=60).encode(
y=alt.Y('NDVI:Q', scale=alt.Scale(domain=(0.4, 0.6))),
color=alt.Color('Year:O', scale=alt.Scale(scheme='magma')),
tooltip=[
alt.Tooltip('Year:O', title='Year'),
alt.Tooltip('PDSI:Q', title='PDSI'),
alt.Tooltip('NDVI:Q', title='NDVI')
])
fit = base.mark_line().encode(
y=alt.Y('Fit:Q'),
color=alt.value('#808080'))
(points + fit).properties(width=600, height=300).interactive()
As you can see, there seems to be some degree of positive correlation between PDSI and NDVI (i.e., as wetness increases, vegetation productivity increases; as wetness decreases, vegetation productivity decreases). Note that some of the greatest outliers are 2016, 2017, 2018 - the three years following recovery from the long drought. It is also important to note that there are many other factors that may influence the NDVI signal that are not being considered here.
Patch-level vegetation mortality
At a regional scale there appears to be a relationship between drought and vegetation productivity. This section will look more closely at effects of drought on vegetation at a patch level, with a specific focus on mortality. Here, a Landsat time series collection is created for the period 1984-present to provide greater temporal context for change at a relatively precise spatial resolution.
Find a point of interest
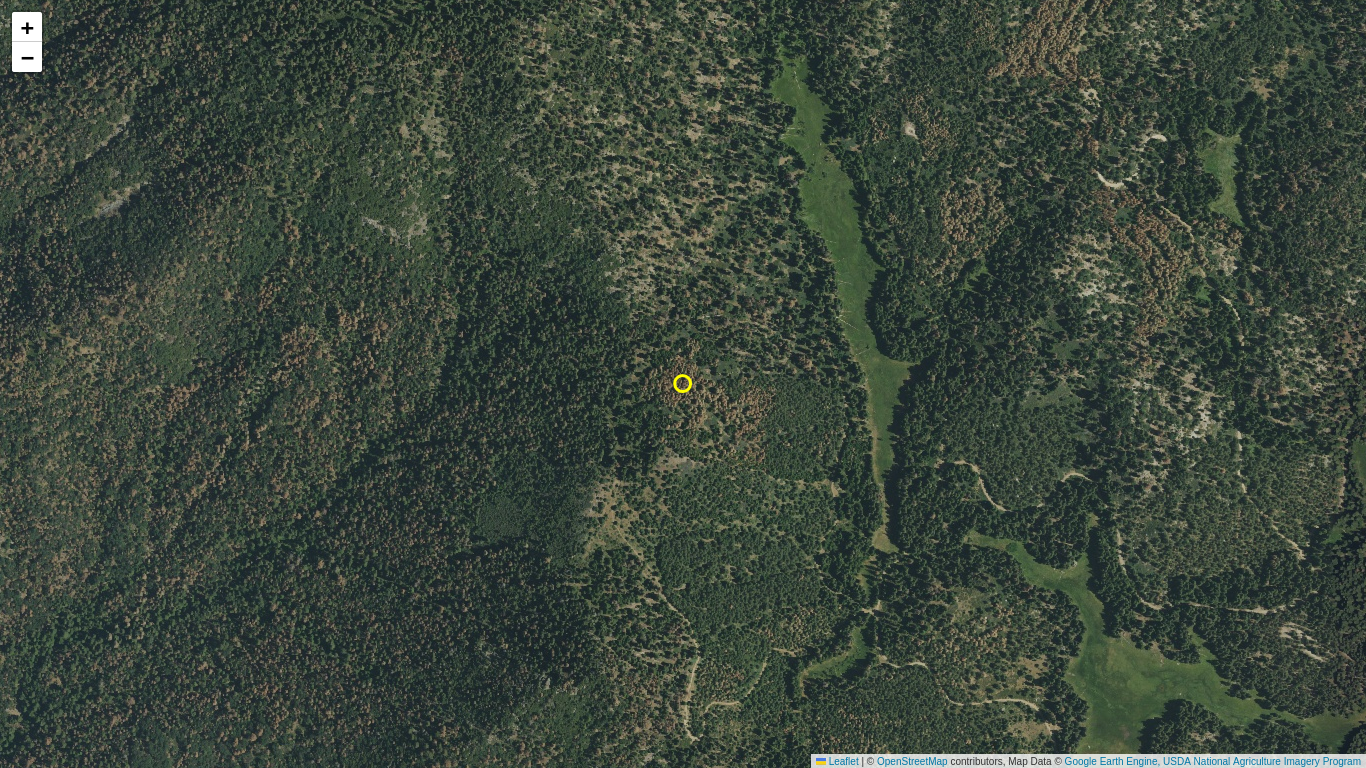
Use aerial imagery from the National Agriculture Imagery Program (NAIP) in an interactive Folium map to identify a location in the Sierra Nevada ecoregion that appears to have patches of dead trees.
- Run the following code block to render an interactive Folium map for a selected NAIP image.
- Zoom and pan around the image to identify a region of recently dead trees (standing silver snags with no fine branches or brown/grey snags with fine branches).
- Click the map to list the latitude and longitude for a patch of interest. Record these values for use in the following section (the example location used in the following section is presented as a yellow point).
# Define a method for displaying Earth Engine image tiles to folium map.
def add_ee_layer(self, ee_image_object, vis_params, name):
map_id_dict = ee.Image(ee_image_object).getMapId(vis_params)
folium.raster_layers.TileLayer(
tiles=map_id_dict['tile_fetcher'].url_format,
attr='Map Data © <a href="https://earthengine.google.com/">Google Earth Engine, USDA National Agriculture Imagery Program</a>',
name=name,
overlay=True,
control=True).add_to(self)
# Add an Earth Engine layer drawing method to folium.
folium.Map.add_ee_layer = add_ee_layer
# Import a NAIP image for the area and date of interest.
naip_img = ee.ImageCollection('USDA/NAIP/DOQQ').filterDate(
'2016-01-01',
'2017-01-01').filterBounds(ee.Geometry.Point([-118.6407, 35.9665])).first()
# Display the NAIP image to the folium map.
m = folium.Map(location=[35.9665, -118.6407], tiles='OpenStreetMap', zoom_start=16)
m.add_ee_layer(naip_img, None, 'NAIP image, 2016')
# Add the point of interest to the map.
folium.Circle(
radius=15,
location=[35.9665, -118.6407],
color='yellow',
fill=False,
).add_to(m)
# Add the AOI to the map.
folium.GeoJson(
aoi.getInfo(),
name='geojson',
style_function=lambda x: {'fillColor': '#00000000', 'color': '#000000'},
).add_to(m)
# Add a lat lon popup.
folium.LatLngPopup().add_to(m)
# Display the map.
display(m)
Prepare Landsat collection
Landsat surface reflectance data need to be prepared before being reduced. The steps below will organize data from multiple sensors into congruent collections where band names are consistent, cloud and cloud shadows have been masked out, and the normalized burn ratio (NBR) transformation is calculated and returned as the image representative (NBR is a good indicator of forest disturbance). Finally, all sensor collections will be merged into a single collection and annual composites calculated based on mean annual NBR using a join.
- Define Landsat observation date window inputs based on NDVI curve plotted previously and set latitude and longitude variables from the map above.
start_day = 224
end_day = 272
latitude = 35.9665
longitude = -118.6407
Note: in your own application it may be necessary to change these values.
Prepare a Landsat surface reflectance collection 1984-present. Those unfamiliar with Landsat might find the following acronym definitions and links helpful.
- OLI (Landsat's Operational Land Imager sensor)
- ETM+ (Landsat's Enhanced Thematic Mapper Plus sensor)
- TM (Landsat's Thematic Mapper sensor)
- CFMask (Landsat USGS surface reflectance mask based on the CFMask algorithm)
- NBR. (Normalized Burn Ratio: a spectral vegetation index)
- Understanding Earth Engine joins
# Make lat. and long. vars an `ee.Geometry.Point`.
point = ee.Geometry.Point([longitude, latitude])
# Define a function to get and rename bands of interest from OLI.
def rename_oli(img):
return (img.select(
ee.List(['SR_B2', 'SR_B3', 'SR_B4', 'SR_B5', 'SR_B6', 'SR_B7']),
ee.List(['Blue', 'Green', 'Red', 'NIR', 'SWIR1', 'SWIR2'])))
# Define a function to get and rename bands of interest from ETM+.
def rename_etm(img):
return (img.select(
ee.List(['SR_B1', 'SR_B2', 'SR_B3', 'SR_B4', 'SR_B5', 'SR_B7']),
ee.List(['Blue', 'Green', 'Red', 'NIR', 'SWIR1', 'SWIR2'])))
# Define a function to scale images and mask out clouds and cloud shadows.
def scale_and_mask(img):
qa_mask = img.select('QA_PIXEL').bitwiseAnd(int('11111', 2)).eq(0)
scaled = img.select('SR_B.').multiply(0.0000275).add(-0.2)
return scaled.updateMask(qa_mask)
# Define a function to add year as an image property.
def set_year(img):
year = ee.Image(img).date().get('year')
return img.set('Year', year)
# Define a function to calculate NBR.
def calc_nbr(img):
return img.normalizedDifference(ee.List(['NIR', 'SWIR2'])).rename('NBR')
# Define a function to prepare OLI images.
def prep_oli(img):
orig = img
img = scale_and_mask(img)
img = rename_oli(img)
img = calc_nbr(img)
img = img.copyProperties(orig, orig.propertyNames())
return set_year(img)
# Define a function to prepare TM/ETM+ images.
def prep_etm(img):
orig = img
img = scale_and_mask(img)
img = rename_etm(img)
img = calc_nbr(img)
img = img.copyProperties(orig, orig.propertyNames())
return set_year(img)
# Import image collections for each Landsat sensor (surface reflectance).
tm_col = ee.ImageCollection('LANDSAT/LT05/C02/T1_L2')
etm_col = ee.ImageCollection('LANDSAT/LE07/C02/T1_L2')
oli_col = ee.ImageCollection('LANDSAT/LC08/C02/T1_L2')
# Filter collections and prepare them for merging.
oli_col = oli_col.filterBounds(point).filter(
ee.Filter.calendarRange(start_day, end_day, 'day_of_year')).map(prep_oli)
etm_col = etm_col.filterBounds(point).filter(
ee.Filter.calendarRange(start_day, end_day, 'day_of_year')).map(prep_etm)
tm_col = tm_col.filterBounds(point).filter(
ee.Filter.calendarRange(start_day, end_day, 'day_of_year')).map(prep_etm)
# Merge the collections.
landsat_col = oli_col.merge(etm_col).merge(tm_col)
# Get a distinct year collection.
distinct_year_col = landsat_col.distinct('Year')
# Define a filter that identifies which images from the complete collection
# match the year from the distinct year collection.
join_filter = ee.Filter.equals(leftField='Year', rightField='Year')
# Define a join.
join = ee.Join.saveAll('year_matches')
# Apply the join and convert the resulting FeatureCollection to an
# ImageCollection.
join_col = ee.ImageCollection(
join.apply(distinct_year_col, landsat_col, join_filter))
# Define a function to apply mean reduction among matching year collections.
def reduce_by_join(img):
year_col = ee.ImageCollection.fromImages(ee.Image(img).get('year_matches'))
return year_col.reduce(ee.Reducer.mean()).rename('NBR').set(
'system:time_start',
ee.Image(img).date().update(month=8, day=1).millis())
# Apply the `reduce_by_join` function to the list of annual images in the
# properties of the join collection.
landsat_col = join_col.map(reduce_by_join)
The result of the above code block is an image collection with as many images as there are years present in the merged Landsat collection. Each image represents the annual mean NBR constrained to observations within the given date window.
Prepare DataFrame
- Create a region reduction function; use
ee.Reducer.first()as the reducer since no spatial aggregation is needed (we are interested in the single pixel that intersects the point). Set the region as the geometry defined by the lat. and long. coordinates identified in the above map. - Apply the function to all images in the time series.
- Filter out features with null computed values.
reduce_landsat = create_reduce_region_function(
geometry=point, reducer=ee.Reducer.first(), scale=30, crs='EPSG:3310')
nbr_stat_fc = ee.FeatureCollection(landsat_col.map(reduce_landsat)).filter(
ee.Filter.notNull(landsat_col.first().bandNames()))
Transfer data from the server to the client.
Note: if the process times out, you'll need to export/import thenbr_stat_fcfeature collection as described in the Optional export section.Convert the Python dictionary to a pandas DataFrame.
Preview the DataFrame and check data types.
nbr_dict = fc_to_dict(nbr_stat_fc).getInfo()
nbr_df = pd.DataFrame(nbr_dict)
display(nbr_df.head())
print(nbr_df.dtypes)
NBR float64 millis int64 system:index object dtype: object
- Add date attribute columns.
- Preview the DataFrame.
nbr_df = add_date_info(nbr_df)
nbr_df.head(5)
Line chart
Display the Landsat NBR time series for the point of interest as a line plot.
alt.Chart(nbr_df).mark_line().encode(
x=alt.X('Timestamp:T', title='Date'),
y='NBR:Q',
tooltip=[
alt.Tooltip('Timestamp:T', title='Date'),
alt.Tooltip('NBR:Q')
]).properties(width=600, height=300).interactive()
As you can see from the above time series of NBR observations, a dramatic decrease in NBR began in 2015, shortly after the severe and extended drought began. The decline continued through 2017, when a minor recovery began. Within the context of the entire time series, it is apparent that the decline is outside of normal inter-annual variability and that the reduction in NBR for this site is quite severe. The lack of major recovery response in NBR in 2017-19 (time of writing) indicates that the event was not ephemeral; the loss of vegetation will have a lasting impact on this site. The corresponding onset of drought and reduction in NBR provides further evidence that there is a relationship between drought and vegetation response in the Sierra Nevada ecoregion.
Past and future climate
The previous data visualizations suggest there is a relationship between drought and vegetation stress and mortality in the Sierra Nevada ecoregion.
This section will look at how climate is projected to change in the future, which can give us a sense for what to expect with regard to drought conditions and speculate about its impact on vegetation.
We'll look at historical and projected temperature and precipitation. Projected data are represented by NEX-DCP30, and historical observations by PRISM.
Future climate
NEX-DCP30 data contain 33 climate models projected to the year 2100 using several scenarios of greenhouse gas concentration pathways (RCP). Here, we'll use the median of all models for RCP 8.5 (the worst case scenario) to look at potential future temperature and precipitation.
Import and prepare collection
- Filter the collection by date and scenario.
- Calculate 'mean' temperature from median min and max among 33 models.
dcp_col = (ee.ImageCollection('NASA/NEX-DCP30_ENSEMBLE_STATS')
.select(['tasmax_median', 'tasmin_median', 'pr_median'])
.filter(
ee.Filter.And(ee.Filter.eq('scenario', 'rcp85'),
ee.Filter.date('2019-01-01', '2070-01-01'))))
def calc_mean_temp(img):
return (img.select('tasmax_median')
.add(img.select('tasmin_median'))
.divide(ee.Image.constant(2.0))
.addBands(img.select('pr_median'))
.rename(['Temp-mean', 'Precip-rate'])
.copyProperties(img, img.propertyNames()))
dcp_col = dcp_col.map(calc_mean_temp)
Prepare DataFrame
- Create a region reduction function.
- Apply the function to all images in the time series.
- Filter out features with null computed values.
reduce_dcp30 = create_reduce_region_function(
geometry=point, reducer=ee.Reducer.first(), scale=5000, crs='EPSG:3310')
dcp_stat_fc = ee.FeatureCollection(dcp_col.map(reduce_dcp30)).filter(
ee.Filter.notNull(dcp_col.first().bandNames()))
- Transfer data from the server to the client. Note: if the process times out, you'll need to export/import the
dcp_stat_fcfeature collection as described in the Optional export section. - Convert the Python dictionary to a pandas DataFrame.
- Preview the DataFrame and check the data types.
dcp_dict = fc_to_dict(dcp_stat_fc).getInfo()
dcp_df = pd.DataFrame(dcp_dict)
display(dcp_df)
print(dcp_df.dtypes)
Precip-rate float64 Temp-mean float64 millis int64 system:index object dtype: object
- Add date attribute columns.
- Preview the DataFrame.
dcp_df = add_date_info(dcp_df)
dcp_df.head(5)
- Convert precipitation rate to mm.
- Convert Kelvin to celsius.
- Add the model name as a column.
- Remove the 'Precip-rate' column.
dcp_df['Precip-mm'] = dcp_df['Precip-rate'] * 86400 * 30
dcp_df['Temp-mean'] = dcp_df['Temp-mean'] - 273.15
dcp_df['Model'] = 'NEX-DCP30'
dcp_df = dcp_df.drop('Precip-rate', axis=1)
dcp_df.head(5)
Past climate
PRISM data are climate datasets for the conterminous United States. Grid cells are interpolated based on station data assimilated from many networks across the country. The datasets used here are monthly averages for precipitation and temperature. They provide a record of historical climate.
Reduce collection and prepare DataFrame
- Import the collection and filter by date.
- Reduce the collection images by region and filter null computed values.
Convert the feature collection to a dictionary and transfer it client-side.
Note: if the process times out, you'll need to export/import theprism_stat_fcfeature collection as described in the Optional export section.Convert the dictionary to a DataFrame.
Preview the DataFrame.
prism_col = (ee.ImageCollection('OREGONSTATE/PRISM/AN81m')
.select(['ppt', 'tmean'])
.filter(ee.Filter.date('1979-01-01', '2019-12-31')))
reduce_prism = create_reduce_region_function(
geometry=point, reducer=ee.Reducer.first(), scale=5000, crs='EPSG:3310')
prism_stat_fc = (ee.FeatureCollection(prism_col.map(reduce_prism))
.filter(ee.Filter.notNull(prism_col.first().bandNames())))
prism_dict = fc_to_dict(prism_stat_fc).getInfo()
prism_df = pd.DataFrame(prism_dict)
display(prism_df)
print(prism_df.dtypes)
/tmpfs/src/tf_docs_env/lib/python3.9/site-packages/ee/deprecation.py:214: DeprecationWarning: Attention required for OREGONSTATE/PRISM/AN81m! You are using a deprecated asset. To make sure your code keeps working, please update it. This dataset has been superseded by OREGONSTATE/PRISM/ANm Learn more: https://developers.google.com/earth-engine/datasets/catalog/OREGONSTATE_PRISM_AN81m warnings.warn(warning, category=DeprecationWarning)
millis int64 ppt float64 system:index object tmean float64 dtype: object
- Add date attribute columns.
- Add model name.
- Rename columns to be consistent with the NEX-DCP30 DataFrame.
- Preview the DataFrame.
prism_df = add_date_info(prism_df)
prism_df['Model'] = 'PRISM'
prism_df = prism_df.rename(columns={'ppt': 'Precip-mm', 'tmean': 'Temp-mean'})
prism_df.head(5)
Combine DataFrames
At this point the PRISM and NEX-DCP30 DataFrames have the same columns, the same units, and are distinguished by unique entries in the 'Model' column. Use the concat function to concatenate these DataFrames into a single DataFrame for plotting together in the same chart.
climate_df = pd.concat([prism_df, dcp_df], sort=True)
climate_df
Charts
Chart the past and future precipitation and temperature together to get a sense for where climate has been and where it is projected to go under RCP 8.5.
Precipitation
base = alt.Chart(climate_df).encode(
x='Year:O',
color='Model')
line = base.mark_line().encode(
y=alt.Y('median(Precip-mm):Q', title='Precipitation (mm/month)'))
band = base.mark_errorband(extent='iqr').encode(
y=alt.Y('Precip-mm:Q', title='Precipitation (mm/month)'))
(band + line).properties(width=600, height=300)
Temperature
line = alt.Chart(climate_df).mark_line().encode(
x='Year:O',
y='median(Temp-mean):Q',
color='Model')
band = alt.Chart(climate_df).mark_errorband(extent='iqr').encode(
x='Year:O',
y=alt.Y('Temp-mean:Q', title='Temperature (°C)'), color='Model')
(band + line).properties(width=600, height=300)
Future climate projections suggest that precipitation will decrease and temperature will increase for the selected point of interest. We can hypothesize, given the RCP 8.5 trajectory, that future conditions will more regularly resemble the 2012-2016 drought, which could lead to the same vegetation reduction response documented here and that more frequent drought events could lead to development of plant communities that are better adapted to low precipitation, high temperature conditions.
 Run in Google Colab
Run in Google Colab
 View source on GitHub
View source on GitHub