April 6th, 2026, Kenneth Philbrick PhD
HealthAI, Google Research
The Imaging Data Commons (IDC) is one of the largest publicly available, de-identified repositories for cancer imaging. In addition to storing imaging, the archive associates and catalogs descriptive metadata and annotations alongside the images. IDC is an invaluable resource for clinicians and researchers, but the sheer scale of the archive presents a challenge: given a novel image, how do you efficiently search across terabytes of pixel imaging to find specific medical or biological features in the archive?
Among its missions, IDC seeks to enable secondary research tasks that are often too resource-intensive to perform on-premises. To support this goal, Google Research has encoded over 65,000 digital pathology whole-slide images from the IDC archive into embeddings. Stored in BigQuery and Google Cloud Storage (GCS), these resources allow researchers to efficiently perform similarity searches across gigapixel images and extract biological insights from the vast IDC archive.
The role of embeddings in generative AI
Machine learning embedding models accelerate the understanding of complex data by transforming images into compact mathematical representations. These 'embeddings' take the form of vectors, lists of floating-point numbers, that functionally encode the concepts within the data. By design, these models cluster related concepts 'close' together in an embedding space, while distinct or unrelated concepts are placed at a greater distance.
The distance between embedding vectors can be quantified using metrics, e.g, Cosine similarity and Euclidean distance. Cosine similarity measures the effective angle between two embedding vectors with values ranging from 1 (identical direction) to -1 (opposite directions). Euclidean distance measures the linear distance between points in the embedding space, with identical vectors having a distance of 0 and the value increasing as the data points become less similar.
Embedding similarity is a core technology that powers many generative AI Retrieval Augmented Generation (RAG) implementations. RAG systems enhance Large Language Model (LLM) prompts by pulling in supplemental data, providing the model with critical information it wasn't originally trained on. Search term embeddings (such as those of images) are typically stored in a vector database. To perform a RAG workflow, an embedding is generated for the specific terms (e.g., text, images, or structured data) within an LLM prompt. The system then retrieves the concepts from the database most closely associated with that embedding and adds them as supplemental context to the prompt. For example, a database could store differential diagnoses linked to medical image embeddings. When a user asks a question about a new image, the system searches the RAG database using that image's embedding and injects the most relevant diagnoses directly into the LLM prompt.
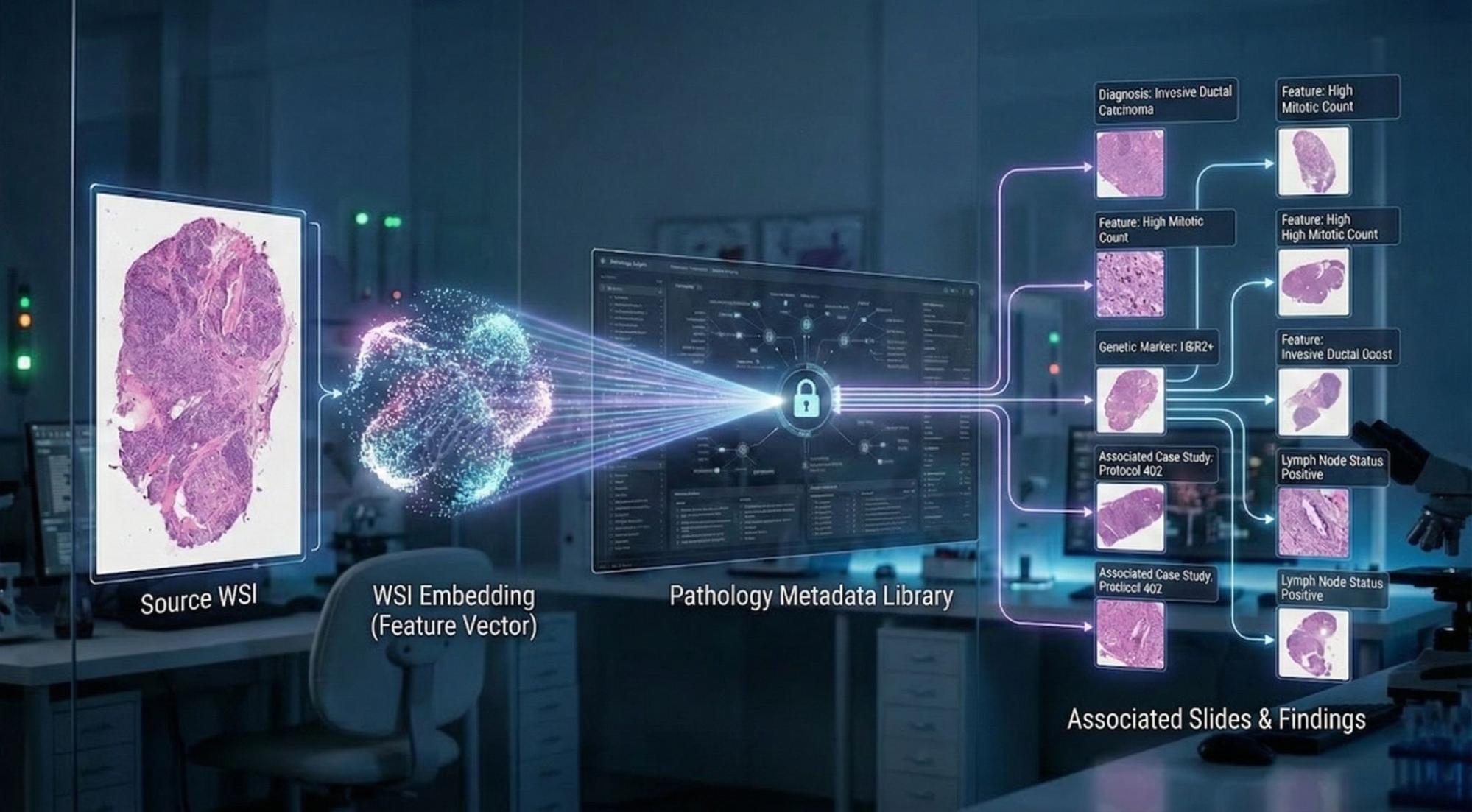
Generating digital pathology embeddings for IDC
To generate digital pathology embeddings at scale, we used resources from Health AI Developer Foundations (HAI-DEF). HAI-DEF offers free open-weight models and open-source tools with an enabling license to help developers build AI-enabled healthcare applications. Specifically, we used the HAI-DEF Path Foundation embedding model to process the IDC digital pathology archive.
Digital pathology images are some of the largest medical images, often reaching gigapixel resolution at the highest magnification. Most machine learning (ML) embedding models cannot process a whole-slide image as a single object; instead, they generate embeddings for "patches" or sub-regions of the image.
To generate embeddings for the IDC digital pathology archive we sampled patches from non-overlapping regions of high magnification tissue imaging (20x - 40x) from 65,802 slides. One Path Foundation embedding (vector of 384 floats) was generated for each sampled patch. On average about 18,000 embeddings were generated per slide to create a total dataset of well over a billion Path Foundation embeddings for the archive as a whole.
Summarizing gigapixel data for search efficiently
To make this vast dataset searchable, we utilized k-means clustering to compress the approximately 18,000 embeddings generated for each slide into a representative set of 500 embeddings. For slides containing fewer than 500 points, we included all generated embeddings. To see an example of this approach you can view the HAI-DEF outlier detector demo, which clusters embeddings of non-tumor tissues for whole-slide imaging from the Camelyon data challenge. The decision to use 500 vectors per slide was determined empirically. Increasing the number would improve the precision of the per-slide representation (preserving less frequent features) but it would increase the total size of the embedding archive. Ultimately, these slide-level embeddings, comprising approximately 33 million total embeddings, were loaded into a BigQuery table and indexed for cosine similarity search.
Performing whole-slide similarity search
With the embedding vectors loaded into BigQuery and indexed, it is now possible to perform whole-slide similarity searches across the entire archive. The process for searching the IDC archive for slide-level similarity is as follows:
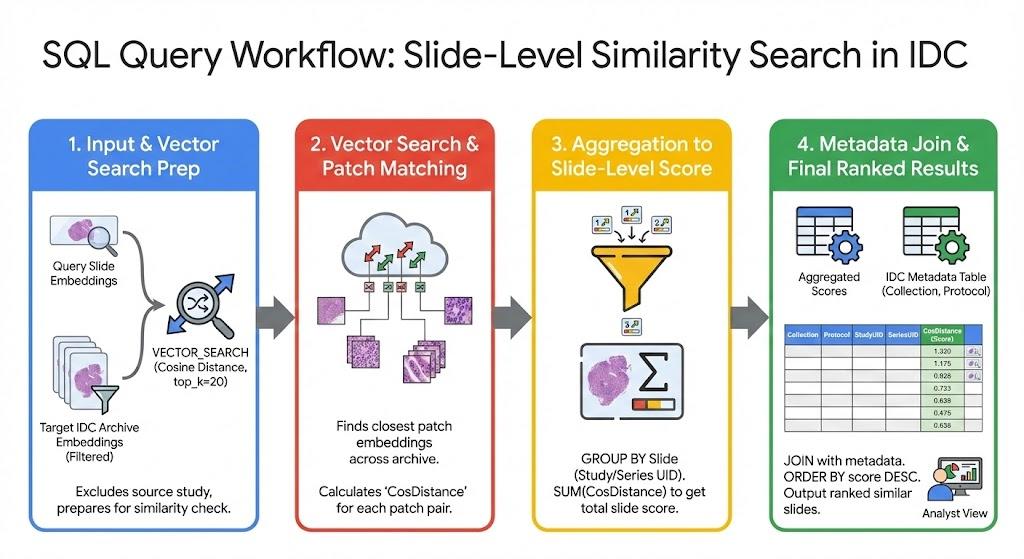
With a novel whole-slide query image at hand, we sample patches across regions of the image. Then we use the HAI-DEF Path Foundation model to encode these patches into embedding vectors and save them to a temporary BigQuery table. In the example shown in the white paper we perform the similarity search using 500 representative embeddings generated from the query image.
We then perform a top-k vector similarity search against the pre-computed IDC embeddings in BigQuery. For each of the 500 patch embeddings representing the query image, we ask BigQuery to return the top K closest matching patches from other whole-slide images from the archive. We determine K empirically. In the example shown in the white paper we return the top 20 matching embeddings for each slide.
The results returned from the BigQuery similarity search identify patch-level embedding similarity. We then aggregate patch-level similarity at the slide level. In the example shown in the white paper we utilize cosine similarity as the embedding similarity metric and aggregate slide-level embedding similarity by summing the top matching patch embeddings for each slide.
We then sort the whole-slide similarity scores and join with additional IDC metadata to identify the specific slides in the archive that best match the query image.
Using BigQuery indexed data for similarity search is highly efficient: archive-level similarity search can be performed in approximately 30 seconds. Whole slide image similarity search and the pre-computed IDC embedding resources are described in greater detail in the white paper.
Resources
Archive-level similarity search is just one example application for these pre-computed IDC imaging embeddings. The HAI-DEF program hosts these indexed embeddings and related metadata in a public GCS bucket and BigQuery dataset and makes them available under the CC BY 4.0 license.
These embeddings were generated using the HAI-DEF Path Foundation embedding model. While this post focuses on pathology, IDC users can apply a similar approach to encode other medical image types. For multi-modal image encoding, we recommend using MedSigLIP.
Pre-computed pathology embeddings and associated metadata using the following repositories:
BigQuery
The dataset contains:
- pathology_embeddings_metadata table: Defines IDC imaging resources available on GCS.
- path_foundation_500_embeddings table: The table used to perform digital pathology image-to-image whole-slide similarity search
GCS bucket
The bucket, (gs://healthai-us/encoded-data/idc/pathology), contains:
- Path Foundation embeddings: Patch-level embeddings generated for each slide.
- Coordinates: The pixel locations where patches were sampled to generate whole-slide image embeddings.
- K-means representations: The reduced representation of each slide's patch embeddings.
- Index mapping: Mapping of patch embeddings to their closest k-means embedding.
- Metadata: Imaging and metadata representing the original imaging and sampled areas.
Date License
The HAI-DEF program hosts these indexed embeddings and related metadata and makes them available to IDC users free of charge under the CC BY 4.0 license.
Next step
Refer to the white paper to see how you can use the pre-computed pathology embeddings to conduct a similar image search on the IDC archive.