April 6th, 2026, Kenneth Philbrick PhD
HealthAI, Google Research
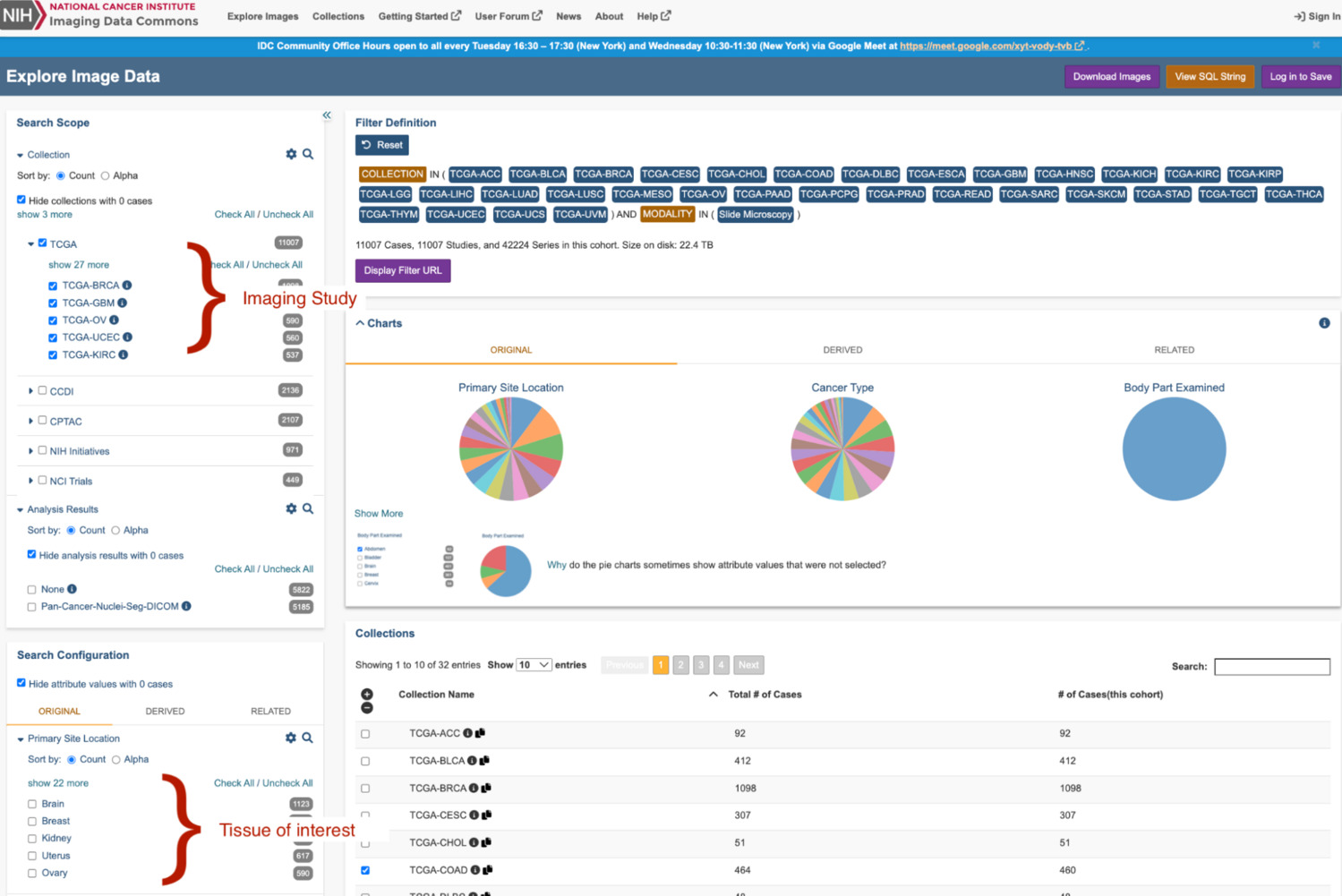
Imaging Data Commons (IDC) is one of the largest publicly available, de-identified, repositories for cancer imaging. The repository is funded by the National Cancer Institute (NCI), an institute of the National Institutes of Health (NIH), a part of the US Department of Health and Human Services. IDC contains imaging for all major medical imaging modalities. Imaging is stored within the archive as DICOM. Imaging and its associated metadata can be searched, visualized through the IDC website, BigQuery, and can be accessed using DICOMweb (IDC, tutorial).
Pathology imaging is considered the gold standard for many cancer diagnoses. To reduce the barrier to the development of novel ML models from the IDC-hosted imaging, we have pre-computed Path Foundation embeddings for the majority of the IDC pathology imaging.
Path Foundation is a patch-based, domain-specific, foundation model. The model was trained using SSL techniques. The model generates embeddings, 384 floats, for input images that are 224 x 224 pixels. The model is available on Hugging Face (open weight) and on Model Garden.
Generation Precomputed Embeddings
IDC version 20, March, 2025, contains about 68 thousand whole slide microscopy images. Precomputed embeddings have been generated for all but a handful of the IDC imaging.
Path Foundation embeddings were computed by sampling patches from the tissue-containing regions of slide imaging. Tissue regions of the slide were determined by thresholding low-magnification imaging, ~2X, and determined by pixels with a HSV value component, 0 < value <= 204; range empirically determined. Non-overlapping 224 x 224 pixel patches were sampled from high-resolution imaging (~20x or greater) and embeddings were computed for the sampled patches.
The number of embeddings generated per-slide range from a few hundred embeddings to well over 100 thousand embeddings. For slides with more than 500 embeddings, embeddings were clustered using scikit-learn MiniBatchKMeans with parameters (n_clusters=500, max_iter=600, n_init=10, batch_size=10240) to generate a reduced representation of the slide's patch embeddings.
K-Means is a clustering technique used here to reduce the large set of embeddings to a smaller representative set of embedding centroids. To see an example of this approach you can view the HAI-DEF outlier detector demo, which clusters embeddings of non-tumor tissues for whole-slide imaging from the Camelyon data challenge to create a model for outlier detection.
For each slide, all generated data is available on Google Cloud Storage (GCS). BigQuery tables were created to join IDC BigQuery tables with GCS resources using joins performed using either the source imaging IDC UUID or DICOM StudyInstanceUID, SeriesInstanceUID, and SOPInstanceUID. Additionally, a BigQuery table has been created that hosts the k-means clustered embeddings. To accelerate search, this data has been indexed using the BigQuery IVF method.
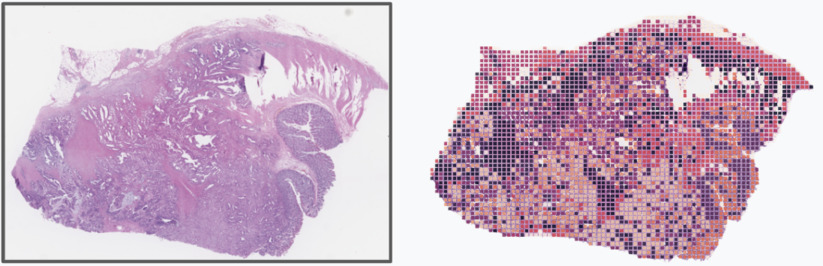
Figure 1: Provides a conceptual illustration of the patch sampling used to generate the pathology embeddings. Each colored square in the image on the right represents a patch sampled for embedding generation. The color of the rectangular patch conceptually represents the index of the k-means generated embedding centroid which was closest to the patch embedding.
Pre-computed Embedding Datasets:
BigQuery Embedding Dataset
GCP Project: hai-cd3-foundations, Dataset: idc_metadata
Dataset License: CC BY 4.0
Resource Table (Table: pathology_embeddings_metadata): Defines mapping between HAI-DEF embedding resources on GCS and Imaging Data Commons imaging.
Search Table (Table: path_foundation_500_embeddings): Contains pre-computed, indexed, digital pathology embeddings for embedding similarity search BigQuery example.
Additional Dataset:
- Imaging Data Commons IDC Archive: IDC generated tables that describe all imaging stored within Imaging Data Commons.
Example: Finding Similar Slides Using BigQuery Vector Search
This example illustrates performing whole slide image similarity search across
IDC imaging using BigQuery vector search. To perform whole slide image
similarity search, embeddings are generated across a search slide and stored
within a BigQuery table. These embeddings define the search set. Vector search
is performed against stored and indexed IDC archive embeddings (N = 32,465,939
embeddings) by performing a series of nested SQL queries using BigQuery. The
embedding similarity search is parameterized to return the top 20 results for
each embedding in the search set. Embedding vector similarity is computed using
cosine similarity.
Per-embedding search results are then aggregated at the slide level to determine
the overall similarity between whole slide images in the IDC archive and the
search set. Slide level aggregation is performed by defining the overall whole
slide image similarity score as the sum of the cosine similarity scores. The sum
of scores works here as an aggregation method because cosine similarity score
metric and embedding similarity are positively correlated. Alternative embedding
search strategies will result in different results. As one example, changing the
distance aggregation metric from sum(CosineSimilarity) to
max(CosineSimilarity) will prioritize the identification of imaging that
closely matches at least one of the search embeddings but otherwise may be quite
different.
Example BigQuery SQL to Perform Similarity Search
SELECT
idc_table.CollectionName AS CollectionName,
idc_table.ProtocolName AS ProtocolName,
embedding_search.StudyInstanceUID AS StudyInstanceUID,
embedding_search.SeriesInstanceUID AS SeriesInstanceUID,
embedding_search.SumCosineSimilarity AS SumCosineSimilarity,
FROM
# Get series descriptive metadata from IDC table.
(
SELECT DISTINCT
collection_name AS CollectionName,
ClinicalTrialProtocolName AS ProtocolName,
StudyInstanceUID,
SeriesInstanceUID
FROM `bigquery-public-data.idc_current.dicom_all`
) AS idc_table,
(
# Get Sum of found search embedding cos distances
# per Unique DICOM (StudyInstanceUID, SeriesInstance UID)imaging.
#
# Cosine Dist
# 1 = vector pointing in same direction
# -1 = vector pointing in opposite direction.
SELECT
StudyInstanceUID, SeriesInstanceUID, sum(CosineSimilarity) AS SumCosineSimilarity
FROM
(
# Return Study instance uid, series instance uid, and
# and distance search embeddings and source..
SELECT
base.StudyInstanceUID AS StudyInstanceUID,
base.SeriesInstanceUID AS SeriesInstanceUID,
distance AS CosineSimilarity
FROM
# Performs a vector search in BQ using COSINE similarity
# Find images which best match search embeddings listed in:
# pathology-cloud-dev.idc_test_us.search_embeddings
VECTOR_SEARCH(
# Search over embeddings excluding any image from DICOM
# that the search embeddings are derived from
(
SELECT *
FROM `hai-cd3-foundations.idc_metadata.path_foundation_500_embeddings`
WHERE
# Exclude any imaging from the same DICOM study that the search
# embeddings were generated. This where clause can be omitted entirely
# if novel imaging is used. If imaging from another IDC study is used
# to perform the search then the StudyInstanceUID should be updated to
# match the values used to define the embedding search (Part 1).
StudyInstanceUID != '2.25.48791557373299768401597362411459861639'
),
'embedding',
TABLE `hai-cd3-foundations.idc_metadata.path_foundation_500_search_example`,
top_k => 20, # Return 20 closest matches for each search embedding
distance_type => 'COSINE', # Embedding search distance metric
options => '{"fraction_lists_to_search": 0.005}')
)
GROUP BY StudyInstanceUID, SeriesInstanceUID
ORDER BY SumCosineSimilarity DESC
LIMIT 100 # Limit results to top 100, closest source embeddings
) AS embedding_search
# Join IDC metadata table with embedding search results.
WHERE
idc_table.StudyInstanceUID = embedding_search.StudyInstanceUID
AND idc_table.SeriesInstanceUID = embedding_search.SeriesInstanceUID
# list results largest first.
ORDER BY SumCosineSimilarity DESC
Example BigQuery Search Results: 
Google Cloud Storage (GCS) Embedding Dataset
Location: gs://healthai-us/encoded-data/idc/pathology
Dataset License: CC BY 4.0
Data for each slide is written to GCS to the following organizational structure:
gs://healthai-us/encoded-data/idc/pathology/{imaging_idc_uuid}/path_foundation/v1/...
Overview of Dataset:
- Metadata (metadata.json): Imaging and metadata representing the original imaging and sampled areas.
Images
- slide_thumbnail.jpeg: Thumbnail of whole slide image.
Example image:
- slide_thumbnail_mask.jpeg: Binary mask projected over thumbnail image illustrating regions of a WSI image sampled for embedding generation.
Example image:
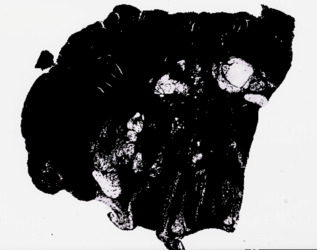
Path Foundation Embeddings (slide_embedding.npy): Patch-level embeddings generated for each slide.
Coordinates (slide_embedding_coord.npy): Location (pixels) in source imaging of left coordinate where each patch was sampled for embedding generation.
K-Means Representations (embeddings_500_cluster_centroid.npy): The k-means reduced representation of each slide's patch embeddings.
Index Mapping (embeddings_500_labels.npy): Maps patch embeddings to their closest k-means embedding.
Detailed description of metadata.json:
| JSON Key | Description |
|---|---|
| instance_uuid | IDC instance uuid |
| instance_hash | IDC generated hash of instance bytes |
| instance_pixeldata_SHA512_hash | Hash of instance pixel data from which embeddings were generated |
| study_instance_uid | DICOM StudyInstanceUID for the imaging used to generate embeddings |
| series_instance_uid | DICOM SeriesInstanceUID for the imaging used to generate embeddings |
| sop_instance_uid | DICOM SOPInstanceUID for the imaging used to generate embeddings |
| instance_patch_embeddings | Sequence of datasets describing patch embedding datasets |
| > model_name | Name of HAI-DEF foundation model used to generate embeddings |
| > model_version | Version of HAI-DEF foundation model used to generate embeddings |
| > version | Embedding dataset version |
| > embedding_count | Number of embeddings generated for the slide |
| > input_image_patch_width_px | Width in pixels of the image patch used to generate embeddings |
| > input_image_patch_height_px | Height in pixels of the image patch used to generate embeddings |
| > patch_embeddings_gs_path | GS path to binary numpy file encoding the slide patch embeddings. The shape of the encoded array is [embedding_count, 384] |
| > patch_coordinates_gs_path | GS path to binary numpy file encoding the slide patch coordinates. The coordinates describe the upper left (X, Y) pixel coordinate of the patch. Coordinates are in reference to the patch source image. The dimensions of the source image are encoded in slide_pyramid_level_width and slide_pyramid_level_height keys. The shape of the encoded array is [embedding_count, 2]. The patch embedding and coordinates arrays are aligned, such that the embeddings described at: patch_embeddings[N] were generated from a patch sampled at patch_coordinates[N]. |
| > thumbnail_patch_sampling_mask_gs_path | GS path to a JPEG image that illustrates where within the image embedding patches were sampled. Image generated by rendering the thumbnail imaging and drawing black squares over the regions sampled by patches. |
| > thumbnail_gs_path | GS path to a JPEG image that illustrates a low resolution image of the tissue imaging. |
| > kmeans_embeddings | Sequence of metadata describing k-means reduced embeddings generated from patch embeddings. |
| >> number_of_centroids | Number of k-means centroids generated. |
| >> centroids_gs_path | GS path to a numpy file encoding the embedding centroids computed by the minibatch_kmeans. The shape of the encoded array is [number of clusters, 384]. |
| >> embedding_centroid_index_gs_path | GS path to a numpy file encoding the index of the embedding centroid that each of the slides embeddings is nearest to using euclidean distance metric. The shape of the encoded array is [embedding_count]. |
Programmatically Accessing the GCS Data using Python
Boilerplate code to convert IDC imaging DICOM StudyInstanceUID and SeriesInstanceUID to IDC instance_uuid.
from google.cloud import bigquery
import google.auth
import requests
def _get_idc_uuid(study_instance_uid: str, series_instance_uid: str) -> str:
"""Utility function to convert DICOM study and series UID to IDC UUID."""
credentials = google.auth.default()[0]
credentials.refresh(google.auth.transport.requests.Request())
# Perform a query.
QUERY = (f'SELECT instance_uuid FROM `hai-cd3-foundations.idc_metadata.pathology_embeddings_metadata` WHERE `StudyInstanceUID` = "{study_instance_uid}" AND `SeriesInstanceUID` = "{series_instance_uid}" LIMIT 1')
client = bigquery.Client(credentials=credentials)
query_job = client.query(QUERY) # API request
results = list(query_job.result())
if not results:
raise ValueError('DICOM instance not found.')
return results[0]['instance_uuid']
Example 1: Read a Slide embedding metadata.json from GCS
import json
import google.cloud.storage as gcs_storage
# Slide is identified by StudyInstanceUID and SeriesInstanceUID
study_instance_uid = '2.25.100000270731860872503075508795340993296'
series_instance_uid = '1.3.6.1.4.1.5962.99.1.1266341113.500336493.1714958324985.4.0'
idc_uuid = _get_idc_uuid(study_instance_uid, series_instance_uid)
client = gcs_storage.Client.create_anonymous_client()
bucket = gcs_storage.Bucket(name='healthai-us', client=client)
blob = bucket.get_blob(f'encoded-data/idc/pathology/{idc_uuid}/path_foundation/v1/metadata.json')
json_str = blob.download_as_string()
print(json.loads(json_str))
Example 2: Read a Slide patch embeddings (numpy file) from GCS
import io
import google.cloud.storage as gcs_storage
import numpy as np
# Slide is identified by StudyInstanceUID and SeriesInstanceUID
study_instance_uid = '2.25.100000270731860872503075508795340993296'
series_instance_uid = '1.3.6.1.4.1.5962.99.1.1266341113.500336493.1714958324985.4.0'
idc_uuid = _get_idc_uuid(study_instance_uid, series_instance_uid)
client = gcs_storage.Client.create_anonymous_client()
bucket = gcs_storage.Bucket(name='healthai-us', client=client)
blob = bucket.get_blob(f'encoded-data/idc/pathology/{idc_uuid}/path_foundation/v1/slide_embedding.npy')
with io.BytesIO() as mem:
blob.download_to_file(mem)
_ = mem.seek(0)
embeddings = np.load(mem)
print(embeddings.shape)
Example 3: Read a Slide Thumbnail Image from GCS
import google.cloud.storage as gcs_storage
# Slide is identified by StudyInstanceUID and SeriesInstanceUID
study_instance_uid = '2.25.100000270731860872503075508795340993296'
series_instance_uid = '1.3.6.1.4.1.5962.99.1.1266341113.500336493.1714958324985.4.0'
idc_uuid = _get_idc_uuid(study_instance_uid, series_instance_uid)
client = gcs_storage.Client.create_anonymous_client()
bucket = gcs_storage.Bucket(name='healthai-us', client=client)
blob = bucket.get_blob(f'encoded-data/idc/pathology/{idc_uuid}/path_foundation/v1/slide_thumbnail.jpeg')
blob.download_to_filename('thumbnail.jpeg')
Generate Pathology Embeddings From DICOM Imaging
The Python library EZ-WSI DICOMweb simplifies generating Path Foundation embeddings from the DICOM imaging. Refer to the IDC documentation for information about how to access digital pathology imaging from the IDC archive using EZ-WSI. The EZ-WSI DICOMweb embedding generation CoLab provides detailed examples that illustrate how to use EZ-WSI DICOMweb to generate Path Foundation embeddings.
Example: Generate a Path Foundation Embedding from a DICOM Whole Slide Image using EZ-WSI DICOMweb
import cv2
from ez_wsi_dicomweb import credential_factory
from ez_wsi_dicomweb import dicom_store
from ez_wsi_dicomweb import patch_embedding
from ez_wsi_dicomweb import patch_embedding_endpoints
# URL for IDC DICOM Store (Requires read access to GCP IDC Store)
# Request for access at:
# https://docs.cloud.google.com/healthcare-api/docs/resources/public-datasets/idc
store_url = 'https://healthcare.googleapis.com/v1/projects/nci-idc-data/locations/us-central1/datasets/idc/dicomStores/idc-store-v20/dicomWeb'
# Connect to IDC DICOM store using application default credentials
ds = dicom_store.DicomStore(store_url, credential_factory.DefaultCredentialFactory())
# Study Instance UID and Series Instance UID of imaging.
study_instance_uid = '2.25.210286627285813284217395096048268203294'
series_instance_uid = '1.3.6.1.4.1.5962.99.1.172459500.862792875.1640849999340.2.0'
slide = ds.get_slide(study_instance_uid, series_instance_uid)
# Load imaging into in memory cache.
cache = slide.init_slide_frame_cache()
# Pathology Embeddings API Natively expects patches to be 224 x 224 pixels
width = 224
height = 224
# Generate a patch slightly offset from the center on the smallest pyramid level.
slide_pyramid_level = list(slide.levels)[-1]
x_coordinate = int((slide_pyramid_level.width - width) / 2) + 100
y_coordinate = int((slide_pyramid_level.height - height) / 2) + 200
patch = slide.get_patch(slide_pyramid_level, x_coordinate, y_coordinate, width, height)
# Defines an endpoint that will generate embeddings using the Google Research Endpoint.
# See linear classifier documentation for examples that illustrate how to
# generate embeddings using the model externalized on Hugging Face.
endpoint = patch_embedding_endpoints.V2PatchEmbeddingEndpoint()
# Generates embedding for a single patch.
# See EZ-WSI embedding documentation for examples which illustrate how
# to generate embeddings in batch for multiple patches or entire slides.
embedding = patch_embedding.get_patch_embedding(endpoint, patch)
# Display embedding
print(embedding)
# optionally retrieve patch pixels to see source imaging for embedding.
image_bytes = patch.image_bytes()
# save image bytes, OpenCV expects image bytes to be in BGR ordering.
cv2.imwrite('test.png', cv2.cvtColor(image_bytes, cv2.COLOR_RGB2BGR))